UniProt provides various resources for those interested in human diseases. Relevant disease information associated with a given protein can be found on the protein entry page in the subsection 'Involvement in disease' of the General annotation section.
One way to find all proteins in UniProt that are associated with a disease is to search for the disease within the 'Human diseases' dataset.
• Select 'Search in': Human diseases from the drop-down next to the Query box
• Type the name of a disease in the Query box
• Click Search
You will be presented with a table of results. For example, let's try searching for 'breast cancer' in the 'human diseases' dataset.
The second hit, 'Breast cancer', matches my query and its description confirms that this is my disease of interest. I see a link to UniProtKB indicating that 10 proteins are involved with this disease and I click on this link to see if it includes any of my proteins of interest.
I look through the list of proteins and see an entry for the BRCA1 human gene, which I am familiar with. I click on the entry link and go to the protein page.
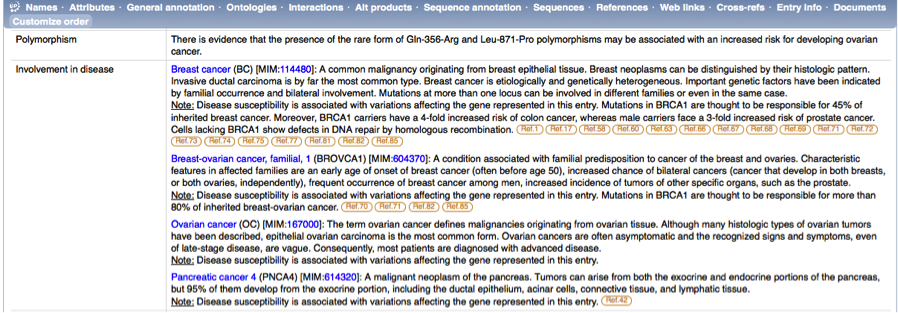

On the protein entry page, I find the 'Involvement in disease' comment line under 'General comments’, which gives me information about the role that this protein plays in breast cancer and links to PubMed references.
I also find a cross-reference to CHEMBL, where I can further investigate all chemical and drug-like compounds that are known to react with this target.
We manually annotate natural variants, including polymorphisms, variations between strains, isolates or cultivars, disease-associated mutations and RNA editing events in UniProtKB entries under the ‘Sequence annotation (Features)’ section. We report the nature of the amino acid change, the name of the variant (or allele), when available, and the effect(s) of the variation on the protein, the cell or the complete organism.
We also provide additional human genetic variation information through FTP downloads. The HUMSAVAR file contains all manually curated human missense variants and the new 1000 Genomes Project variants file contains a catalogue of novel Single Nucleotide Variants (SVNs or SNPs) from the 1000 Genomes Project for both UniProtKB/Swiss-Prot and UniProtKB/TrEMBL sequences. Both files can be downloaded at UniProt's FTP site.


